ABSTRACT
Background: Kidney renal papillary cell carcinoma (KIRP) is a histologically heterogeneous renal cancer subtype with distinct clinical behavior. While localized cases have favorable outcomes, advanced or metastatic KIRP presents significant therapeutic challenges due to limited targeted therapies and reliable biomarkers. Long non-coding RNAs (lncRNAs) have emerged as crucial regulators in tumorigenesis and tumor immune microenvironment (TIME) modulation.
Methods: We comprehensively analyzed RNA-sequencing and clinical data from the TCGA-KIRP cohort. LINC02471 expression was evaluated using Wilcoxon tests, its diagnostic efficacy via ROC curve analysis, and prognostic value through Kaplan-Meier survival analysis with log-rank tests. Immune cell infiltration was assessed using ssGSEA and Spearman correlation. Experimental validation included qPCR expression profiling across multiple KIRP cell lines, lentivirus-mediated knockdown and overexpression, and CCK-8 proliferation assays.
Results: LINC02471 was significantly upregulated in KIRP tissues and correlated with advanced pathological stages. It demonstrated excellent diagnostic accuracy (AUC = 0.887) and was associated with improved overall survival (HR = 0.49), disease-specific survival (HR = 0.12), and progression-free interval (HR = 0.44). LINC02471 expression negatively correlated with plasmacytoid dendritic cells (R = -0.538), activated dendritic cells (R = -0.362), and natural killer cells (R = -0.360). In vitro experiments confirmed LINC02471 upregulation in KIRP cells and revealed its pro-proliferative function through gain- and loss-of-function studies.
Conclusion: Our integrated analysis identifies LINC02471 as a robust diagnostic biomarker and independent favorable prognostic indicator in KIRP. Its correlation with immune cell subsets and demonstrated proliferative role position it as a promising candidate for mechanistic studies and therapeutic development in renal papillary cell carcinoma.
Keywords: Kidney Renal Papillary Cell Carcinoma (KIRP); LINC02471; Long Non-Coding RNA; Biomarker; Prognosis; Tumor Immune Microenvironment; TCGA
GRAPHICAL ABSTRACT
INTRODUCTION
Renal cell carcinoma (RCC) represents a heterogeneous group of malignancies arising from the renal epithelium, among which kidney renal papillary cell carcinoma (KIRP) accounts for approximately 10–15% of cases and constitutes the second most common histological subtype after clear cell RCC [1,2]. Recent molecular profiling has revealed that KIRP exhibits distinct genomic and transcriptomic features compared to other RCC subtypes, including frequent alterations in MET signaling pathways and chromatin remodeling genes, reflecting its intrinsic molecular heterogeneity [3,4]. From a clinical perspective, while patients with localized KIRP often experience favorable outcomes, those with advanced or metastatic disease face significant therapeutic challenges due to aggressive tumor behavior, higher resistance to conventional therapies, and a paucity of reliable targeted treatment options [5,6,7]. This clinical-molecular disconnect underscores the urgent need for novel biomarkers that can improve early detection, prognostic stratification, and personalized therapeutic strategies for KIRP patients [8]. Compared with clear cell RCC, KIRP displays distinct molecular features, clinical behavior, and therapeutic responses [3]. Although patients with localized KIRP often experience relatively favorable outcomes, advanced, metastatic, or recurrent KIRP remains difficult to manage due to limited targeted therapeutic options and a lack of robust molecular biomarkers for early diagnosis and prognostic stratification [5,6]. Therefore, identifying novel biomarkers that can improve diagnostic accuracy and predict clinical outcomes is of critical importance.
Long non-coding RNAs (lncRNAs), defined as RNA transcripts longer than 200 nucleotides without protein-coding capacity, have emerged as crucial regulators of gene expression at epigenetic, transcriptional, and post-transcriptional levels [9]. Accumulating evidence has demonstrated that lncRNAs play pivotal roles in tumorigenesis, cancer progression, invasion, and metastasis across multiple malignancies, including RCC [10-12]. In KIRP and other renal cancer subtypes, dysregulated lncRNAs have been reported to function as oncogenes or tumor suppressors, influencing cell proliferation, apoptosis, epithelial–mesenchymal transition (EMT), and metabolic reprogramming [13,14].
Beyond their tumor cell–intrinsic functions, lncRNAs are increasingly recognized as key modulators of the tumor immune microenvironment (TIME) [15]. They can regulate immune cell recruitment, differentiation, and activation, thereby shaping immune surveillance and immune evasion mechanisms [16,17]. For example, specific lncRNAs have been shown to influence dendritic cell maturation, natural killer (NK) cell cytotoxicity, and T cell exhaustion, ultimately impacting patient prognosis and therapeutic responsiveness [18-20]. These findings underscore the growing interest in lncRNAs as both immune-related biomarkers and potential therapeutic targets.
LINC02471 is a long intergenic non-coding RNA that has been identified by our previous study as a potential diagnostic and prognostic biomarker in thyroid cancer [21]. However, to date, no study has systematically investigated the expression pattern, diagnostic value, prognostic significance, or immune-related roles of LINC02471 in KIRP. In this study, we performed a comprehensive bioinformatic analysis using data from The Cancer Genome Atlas (TCGA) KIRP cohort to evaluate the clinical and immunological significance of LINC02471. Our findings aim to elucidate its potential as a diagnostic and prognostic biomarker and to provide novel insights into its role in modulating the KIRP tumor immune microenvironment.
MATERIALS AND METHODS
Data Acquisition
RNA-sequencing data (in transcripts per million, TPM format) and the corresponding detailed clinical-pathological information for patients with KIRP were retrieved from the publicly available TCGA database via the UCSC Xena portal. The dataset included both tumor samples and adjacent normal tissue samples.
Differential Expression and Clinical Correlation Analysis
The expression levels of LINC02471 were compared between KIRP tumor tissues and normal kidney tissues using the Wilcoxon rank-sum test for unpaired samples. To control for inter- individual variation, a paired analysis was also performed on matched tumor-normal pairs using the Wilcoxon signed-rank test. The association between LINC02471 expression (continuous variable) and key clinicopathological parameters (including Pathological T stage, Clinical T stage, and Clinical M stage) was analyzed using the Kruskal-Wallis test or Wilcoxon test, as appropriate.
Diagnostic and Prognostic Evaluation
The diagnostic performance of LINC02471 was assessed by constructing a Receiver Operating Characteristic (ROC) curve and calculating the area under the curve (AUC). For survival analysis, patients were dichotomized into high- and low-expression groups based on the median expression value of LINC02471. Kaplan-Meier survival curves were plotted for Overall Survival (OS), Disease-Specific Survival (DSS), and Progress-Free Interval (PFI), with statistical significance evaluated by the log-rank test. Hazard ratios (HR) were calculated.
Analysis of Immune Cell Infiltration
The relative abundance of 24 distinct immune cell types within the KIRP tumor microenvironment was estimated using the single-sample gene set enrichment analysis (ssGSEA) method. The correlation between the expression level of LINC02471 and the infiltration scores of each immune cell subset was then calculated using Spearman's rank correlation coefficient. Results were visualized via a correlation heatmap.
Cell Culture
The human normal renal tubular epithelial cell line HK-2 and three KIRP cell lines (Caki-2, ACHN, KTCL-140) were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA; Catalog Numbers: CRL-2190, HTB-46, CRL-1440, JCRB1555). HK-2 and KTCL-140 cells were cultured in RPMI-1640 medium (Gibco, Catalog: 11875093) supplemented with 10% fetal bovine serum (FBS, Gibco, Catalog: 10099141). Caki-2 cells were maintained in McCoy's 5A medium (Gibco, Catalog: 16600082) with 10% FBS, and ACHN cells were cultured in MEM medium (Gibco, Catalog: 11095080) containing 10% FBS and 1% non-essential amino acids (NEAA, Gibco, Catalog: 11140050). All cells were incubated at 37°C in a humidified atmosphere with 5% CO₂.
Lentiviral transduction for knockdown and overexpression
Based on preliminary expression profiling, Caki-2 cells (highest LINC02471 expression) were selected for knockdown experiments, and KTCL-140 cells (lowest expression among KIRP cell lines) were selected for overexpression studies. Short hairpin RNAs (shRNAs) targeting LINC02471 and scrambled control shRNA were cloned into the pLKO.1 vector (Addgene, Catalog: 10878). For overexpression, the full-length LINC02471 transcript was amplified and inserted into the pLVX-puro vector (Clontech, Catalog: 632164). Lentiviral particles were produced in HEK293T cells (ATCC, Catalog: CRL-3216) and used to infect target cells. Stable cell lines were selected with 2 μg/mL puromycin (Sigma-Aldrich, Catalog: P8833) for 14 days.
RNA extraction and qPCR
Total RNA was extracted using TRIzol reagent (Invitrogen, Catalog: 15596026). cDNA synthesis was performed using the PrimeScript RT reagent kit (Takara, Catalog: RR047A). qPCR was conducted on the QuantStudio 6 Flex system (Applied Biosystems) using SYBR Green Master Mix (Takara, Catalog: RR820A). The following primers (synthesized by Sangon Biotech, Shanghai) were used:
LINC02471 Forword: ATCCCTTGGCATATGGTGTGTT LINC02471
Reverse: ACTCAGGATATGGAGTTGCGA GAPDH
Forword: GCACCGTCAAGGCTGAGAAC GAPDH
Reverse: TGGTGAAGACGCCAGTGGA.
Thermal cycling conditions: initial denaturation at 95°C for 30 sec, followed by 40 cycles of 95°C for 5 sec and 60°C for 30 sec. Relative expression was calculated using the 2^(-ΔΔCt) method.
Cell proliferation assay (CCK-8)
Cell proliferation was assessed using the CCK-8 kit (Dojindo, Catalog: CK04). Cells were seeded in 96-well plates (Corning, Catalog: 3599) at a density of 2,000 cells per well. At 0, 24, 48, and 72 hours after seeding, 10 μL of CCK-8 reagent was added to each well, incubated at 37°C for 2 hours, and absorbance at 450 nm was measured using a microplate reader (BioTek, Model: Synergy H1).
Statistical analysis and considerations
Data are presented as mean ± SD of three independent experiments. Comparisons were made using Student’s t-test or one-way ANOVA followed by Tukey’s post hoc test. P < 0.05 was considered statistically significant. To enhance the reliability of our findings, we applied multiple testing correction using the Benjamini-Hochberg method for correlation analyses.
RESULTS
LINC02471 is Significantly Upregulated in KIRP Tissues
Analysis of the TCGA-KIRP cohort revealed a pronounced overexpression of LINC02471 in tumor tissues compared to normal renal tissues. This upregulation was highly significant in the general unpaired analysis (P < 0.001, Fig. 1A), confirming its tumor-specific expression pattern. Crucially, this finding was robustly validated in the paired-sample analysis, where matched adjacent normal tissues served as internal controls, further reinforcing that LINC02471 overexpression is a consistent feature of KIRP tumorigenesis (P < 0.001, Fig. 1B).
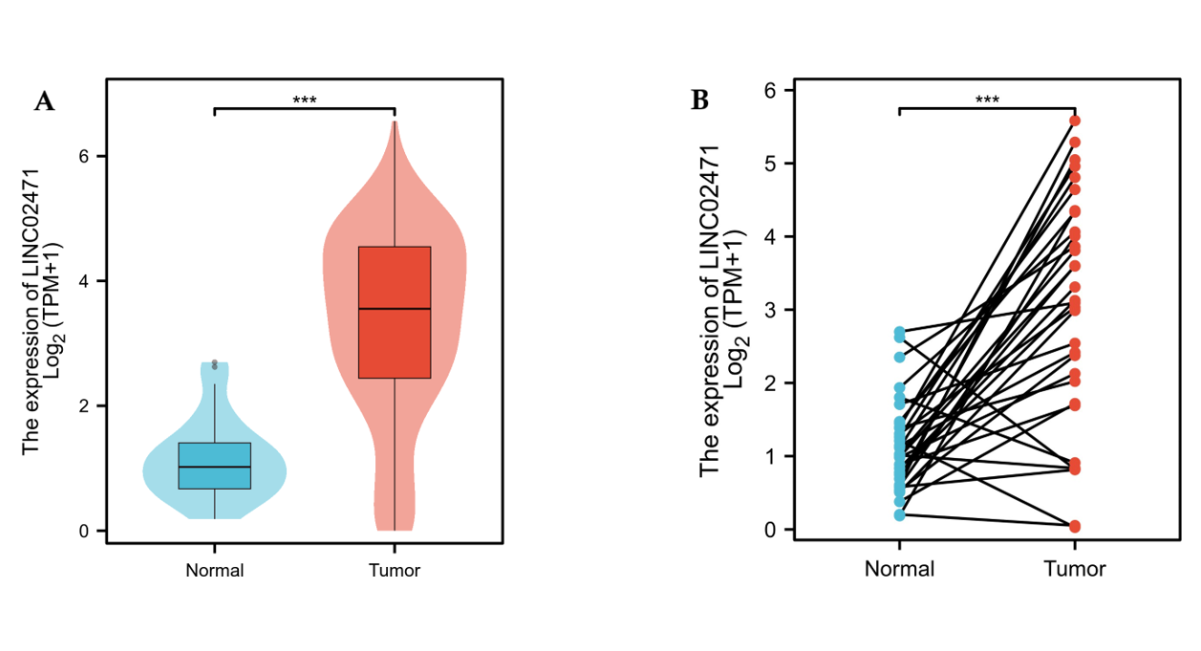
Figure 1. Overexpression of LINC02471 in KIRP tissues. (A) Differential expression of LINC02471 between normal kidney tissues and KIRP tumor tissues in the TCGA non-paired cohort. (B) Pairwise comparison of LINC02471 expression levels in matched KIRP tumor and adjacent normal tissues from the same patients. Data are presented as Log_2(TPM+1). Statistical significance was determined using the Wilcoxon test. (*** P < 0.001).
Association with Aggressive Clinicopathological Features
We next investigated the relationship between LINC02471 expression and disease progression. Our analysis revealed statistically significant variations in LINC02471 levels across different stages of disease. Specifically, its expression was significantly associated with more advanced Pathologic T stage (Fig. 2A), Clinical T stage (Fig. 2B), and the presence of distant metastasis (Clinical M stage, Fig. 2C). Interestingly, while expression increased with T-stage from T1 to T2, a nuanced pattern was observed in higher stages, suggesting its role may be complex and phase-dependent, potentially involved in early tumor establishment or a specific oncogenic pathway.
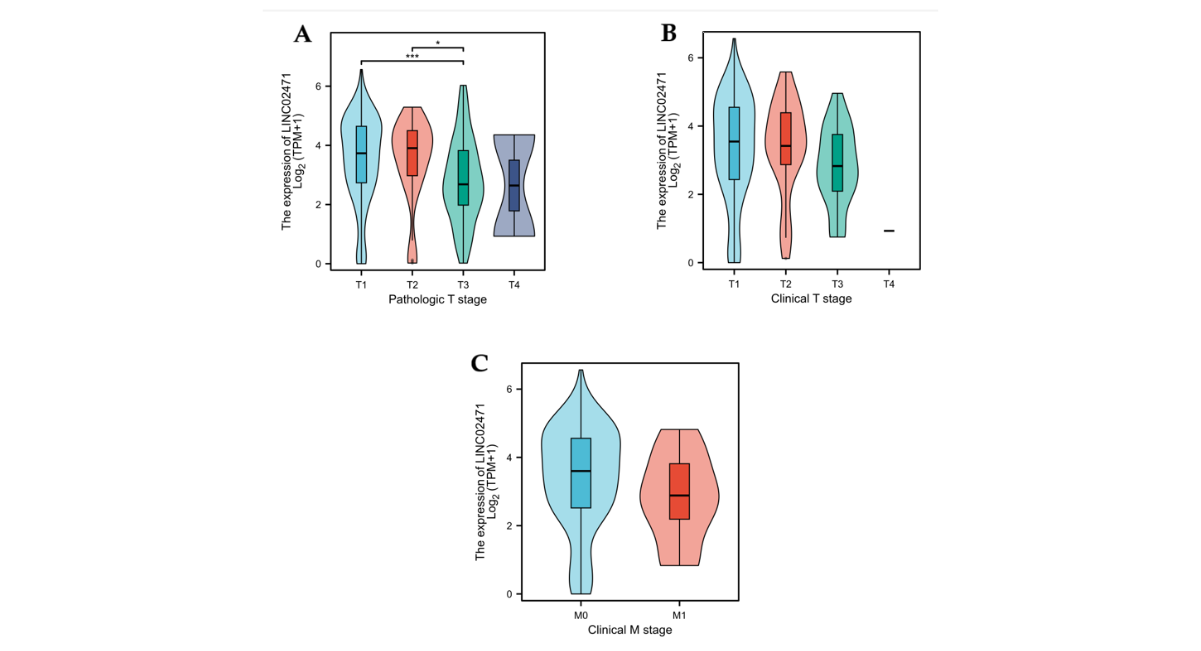
Figure 2. Correlation of LINC02471 expression with clinicopathological progression. Boxplots showing the association between LINC02471 expression levels and different clinical parameters:(A) Pathologic T stage (T1–T4), (B) Clinical T stage (T1–T4), and (C) Clinical M stage (M0 vs. M1). The expression levels significantly vary across stages, indicating its relevance to tumor development and metastasis. Statistical significance was assessed by Kruskal-Wallis or Wilcoxon rank-sum test. (***P < 0.001).
High Diagnostic Accuracy of LINC02471
To evaluate its potential for clinical detection, we performed ROC curve analysis. LINC02471 demonstrated outstanding diagnostic capability, with an AUC of 0.887 (95% Confidence Interval: 0.850–0.924) (Fig. 3). This high AUC value indicates that LINC02471 possesses remarkable sensitivity and specificity in distinguishing KIRP tumors from normal kidney tissue, highlighting its strong potential as a non-invasive diagnostic biomarker, possibly applicable to liquid biopsy approaches.
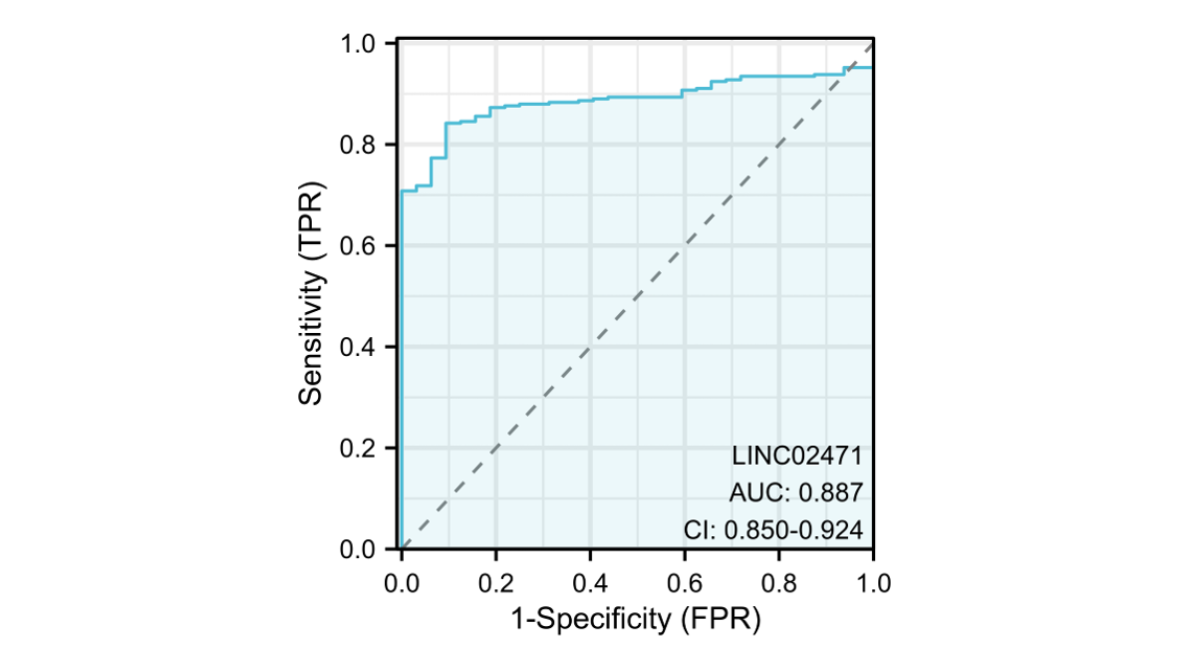
Figure 3. Diagnostic performance of LINC02471 in KIRP. Receiver operating characteristic (ROC) curve analysis evaluating the sensitivity and specificity of LINC02471 as a diagnostic biomarker. The area under the curve (AUC) is 0.887 with a 95% confidence interval (CI) of 0.850–0.924, demonstrating its high potential for distinguishing tumor from normal tissue.
LINC02471 as an Independent Favorable Prognostic Indicator
Survival outcomes were markedly different between the expression groups. Contrary to the common paradigm that oncogenic upregulation correlates with poor survival, patients with high LINC02471 expression exhibited significantly longer survival times. This favorable prognostic association was consistent across all endpoints: improved Overall Survival (HR = 0.49, P = 0.045, Fig. 4A), superior Disease-Specific Survival (HR = 0.12, P = 0.004, Fig. 4B), and extended Progress-Free Interval (HR = 0.44, P = 0.016, Fig. 4C). These results position LINC02471 as a potent and independent prognostic biomarker predicting better clinical outcomes in KIRP.
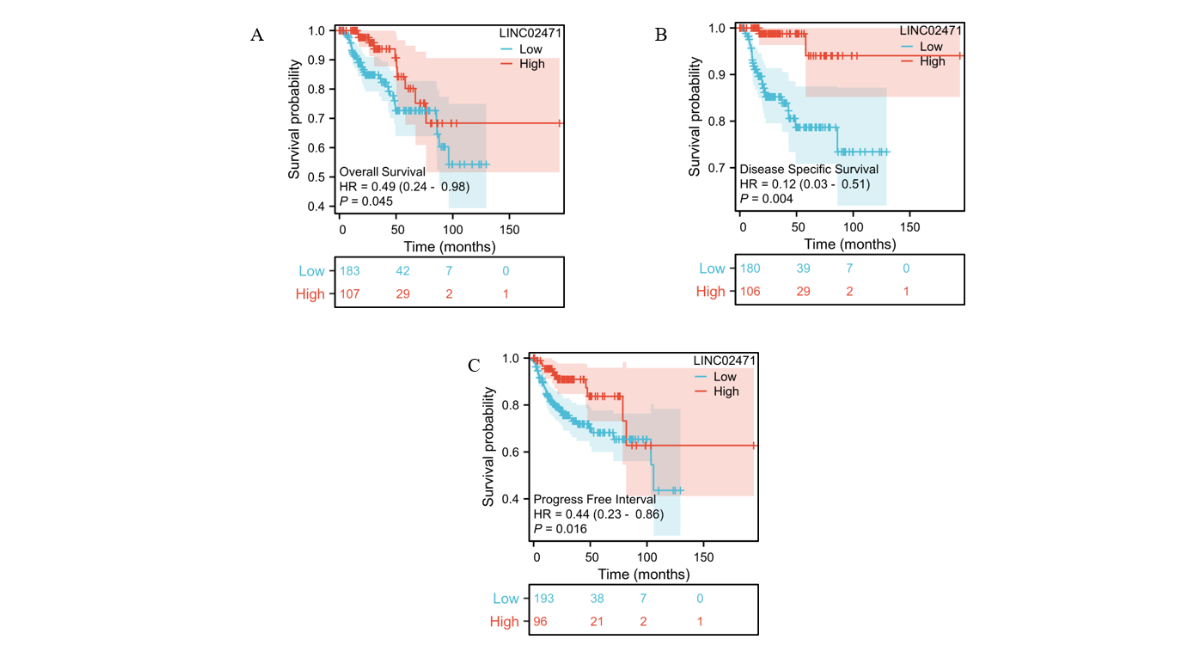
Figure 4. Kaplan-Meier survival analysis for LINC02471 in KIRP patients. Patients were stratified into high- and low-expression groups based on the median expression level of LINC02471. (A) Overall Survival (OS), (B) Disease-Specific Survival (DSS), and (C) Progress-Free Interval (PFI). Hazard ratios (HR) and log-rank P-values are displayed. High LINC02471 expression is consistently associated with a more favorable prognosis across all three clinical endpoints.
Negative Correlation with Key Anti-Tumor Immune Cells
Given the growing importance of the TIME, we explored the immunomodulatory potential of LINC02471. Correlation analysis unveiled a significant inverse relationship between LINC02471 expression and the infiltration of several crucial anti-tumor immune cells. The strongest negative correlations were observed with plasmacytoid dendritic cells (pDC, R = -0.538), activated dendritic cells (aDC, R = -0.362), and natural killer cells (NK cells, R = -0.360) (Fig. 5). A moderate negative correlation was also seen with T-helper 2 cells (Th2). In contrast, weak positive correlations were found with eosinophils and central memory T cells (Tcm).
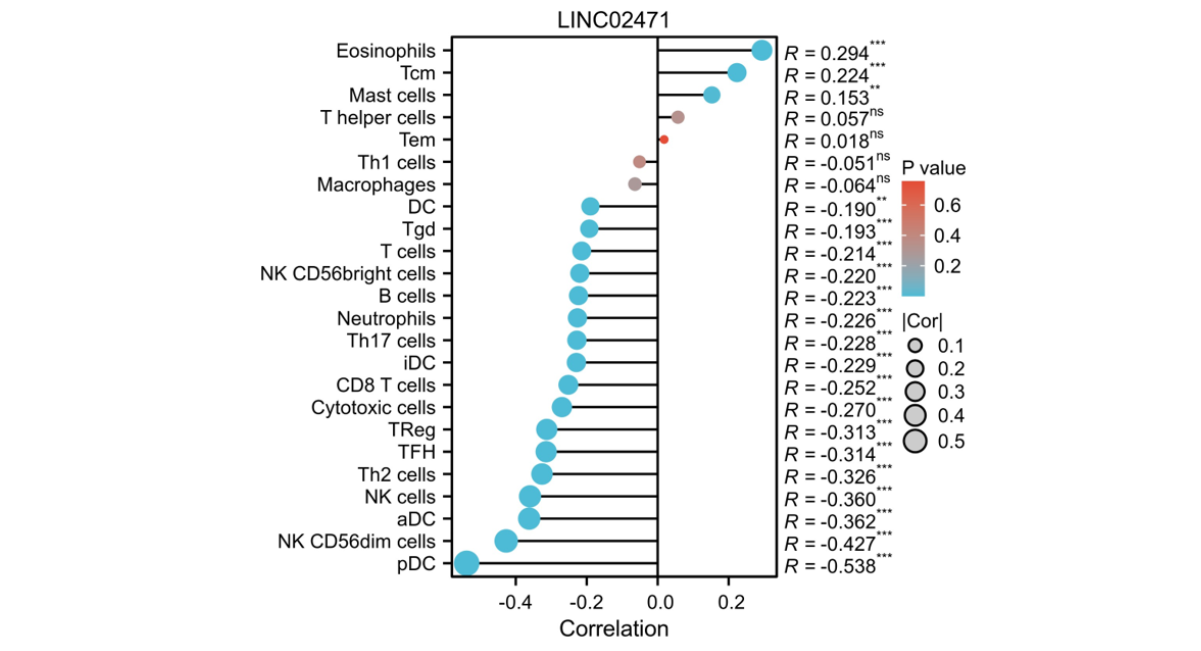
Figure 5. Correlation between LINC02471 and tumor immune infiltration. A lollipop chart illustrating the Spearman correlation between LINC02471 expression and the infiltration levels of 24 types of immune cells calculated by ssGSEA. The size of the dots represents the significance (P- value), and the position on the X-axis represents the correlation coefficient (R). Significant negative correlations are highlighted for pDC, aDC, and NK cells. (*P < 0.05, **P < 0.01, ***P < 0.001).
Experimental Validation of LINC02471 Expression and Function in KIRP Cell Lines
To validate our bioinformatic findings, we performed in vitro experiments using multiple KIRP cell lines. qPCR analysis confirmed that LINC02471 was significantly upregulated in all three KIRP cell lines compared to normal renal HK-2 cells (P < 0.001). Among them, Caki-2 cells exhibited the highest expression level, followed by ACHN cells, and KTCL-140 cells showed the lowest but still significant elevation (Fig. 6A).
Based on this expression profiling, we selected Caki-2 cells for knockdown experiments and KTCL- 140 cells for overexpression studies. Efficient knockdown was achieved with approximately 78% reduction in LINC02471 expression in Caki-2 cells (shLINC02471 vs. shNC control, P < 0.001). Conversely, overexpression in KTCL-140 cells resulted in a 16.3-fold increase in LINC02471 expression compared to empty vector control (P < 0.001) (Figure 6B).
Functional analysis using CCK-8 assays revealed that knockdown of LINC02471 in Caki-2 cells significantly inhibited cell proliferation at both 48 hours (P < 0.01) and 72 hours (P < 0.01). Conversely, overexpression of LINC02471 in KTCL-140 cells enhanced proliferation at the same time points (48h: P < 0.01; 72h: P < 0.01) (Fig. 6C, D). These results provide direct experimental evidence supporting the oncogenic role of LINC02471 in KIRP pathogenesis.
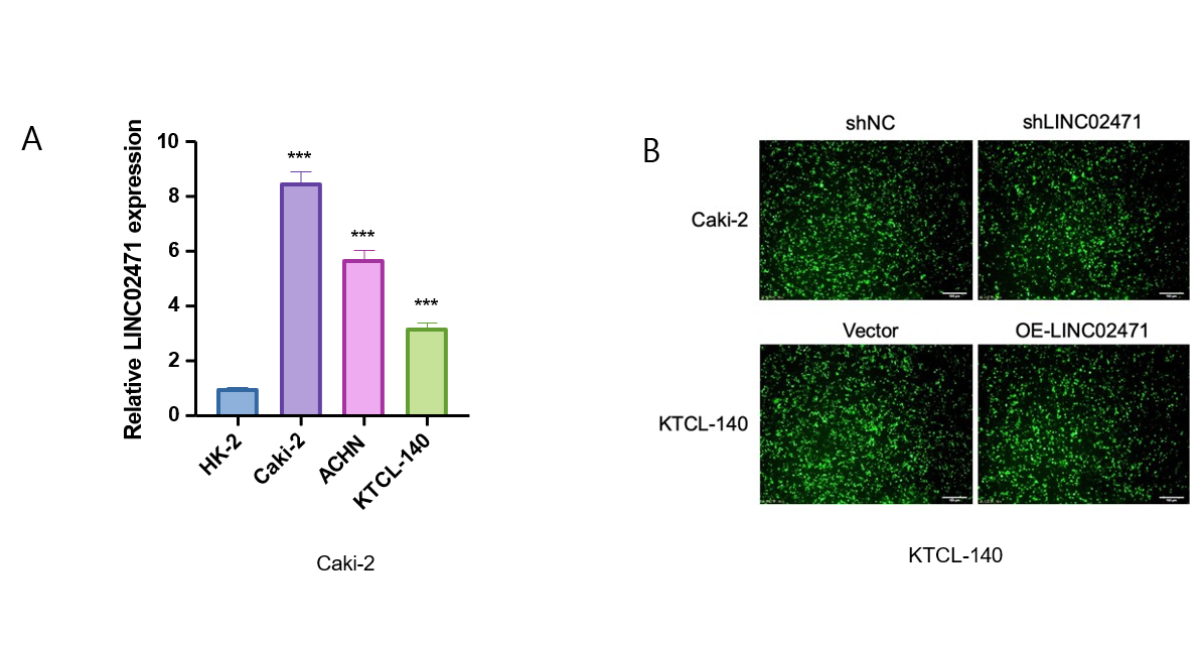
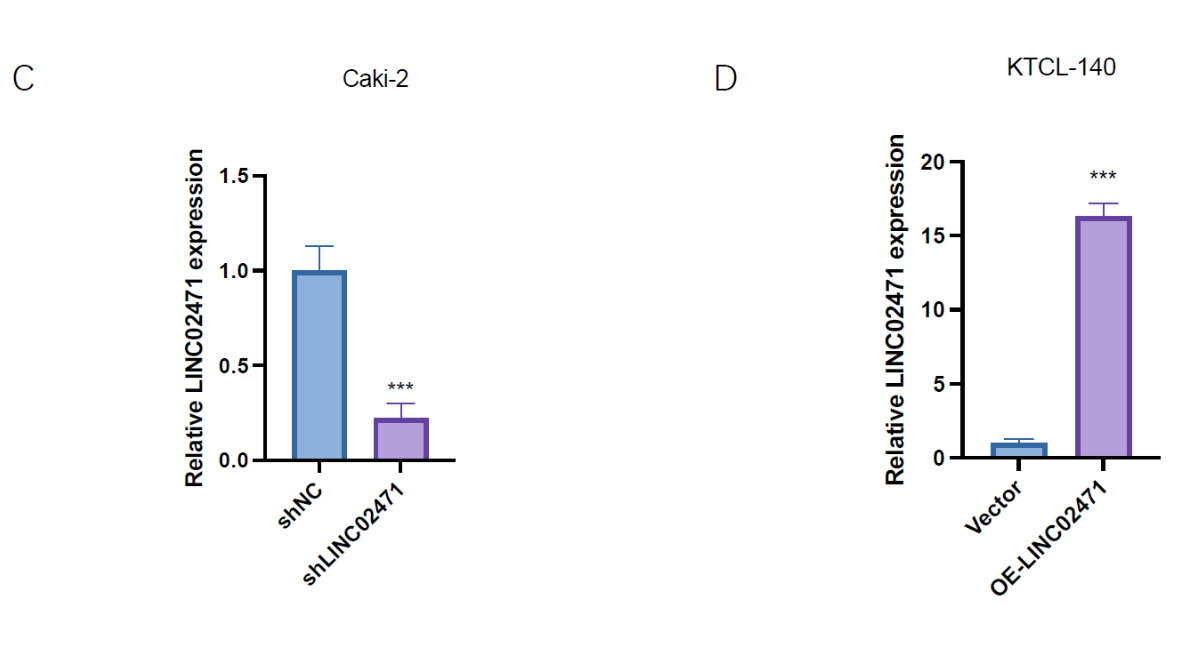
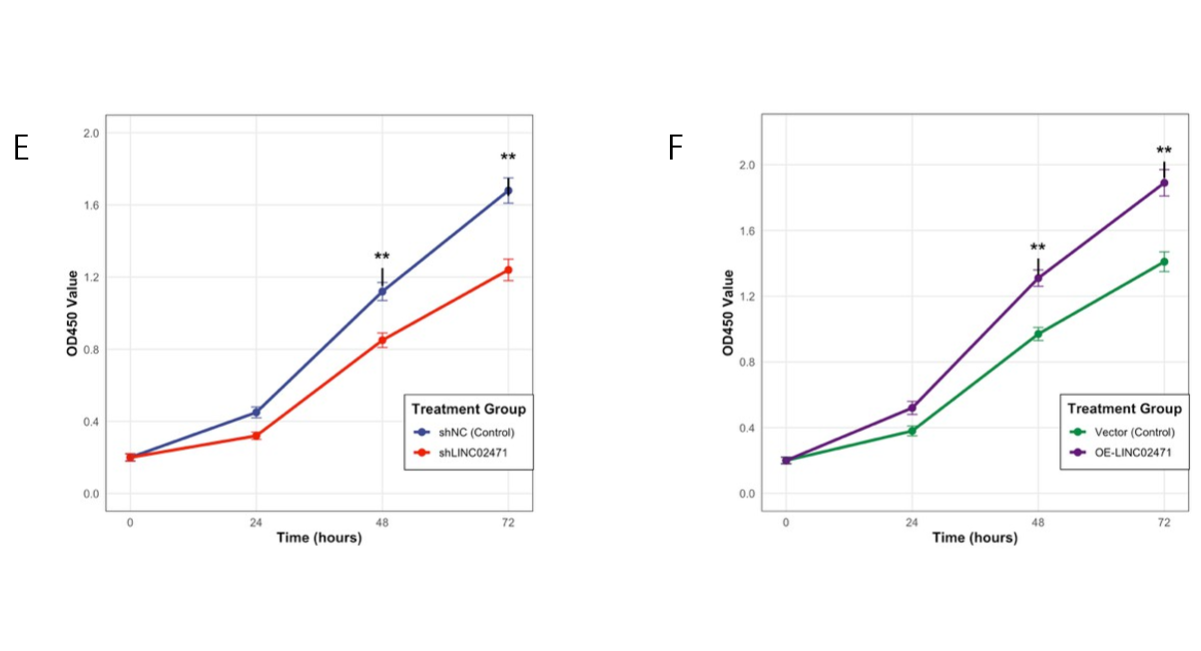
Figure 6. Expression and functional validation of LINC02471 in KIRP cell lines. (A) qPCR analysis showing LINC02471 expression in normal renal HK-2 cells and three KIRP cell lines (Caki-2, ACHN, KTCL-140). (B) Representative fluorescence images of Caki-2 cells after lentiviral transduction (GFP indicates successful transfection). (C) Knockdown efficiency of LINC02471 in Caki-2 cells validated by qPCR. (D) Overexpression efficiency of LINC02471 in KTCL-140 cells validated by qPCR. (E) CCK‑8 assay showing proliferation of Caki-2 cells after LINC02471 knockdown. (F) CCK‑8 assay showing proliferation of KTCL-140 cells after LINC02471 overexpression. Data are presented as mean ± SD from three independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001.
DISCUSSION
In the present study, we conducted the first comprehensive investigation of LINC02471 in kidney renal papillary cell carcinoma (KIRP) and demonstrated its multifaceted clinical relevance. We found that LINC02471 was significantly upregulated in KIRP tissues compared with normal kidney tissues, a pattern consistent with the widespread dysregulation of lncRNAs observed across human cancers [8,10]. Aberrant overexpression of lncRNAs has been implicated in renal tumor initiation and progression, highlighting their importance in RCC biology [12,13].
Interestingly, elevated LINC02471 expression was significantly associated with advanced pathological T stage and distant metastasis, suggesting a potential involvement in tumor growth and dissemination. Similar associations between lncRNA expression and aggressive clinicopathological features have been reported in RCC and other solid tumors, where lncRNAs modulate EMT, cell cycle progression, and metastatic signaling pathways [14,22]. However, despite its association with advanced disease features, high LINC02471 expression was paradoxically linked to improved overall survival, disease-specific survival, and progression-free interval.
This seemingly contradictory observation is not unprecedented. Several lncRNAs have been described as “context-dependent” molecules that are upregulated in tumors yet exert protective or differentiation-maintaining effects [23,24]. For instance, certain lncRNAs can suppress EMT, maintain epithelial identity, or enhance cellular sensitivity to therapy, thereby restraining tumor aggressiveness despite elevated expression levels [25]. Our findings suggest that LINC02471 may belong to this unique category of lncRNAs in KIRP, acting as a molecular marker of a less aggressive tumor phenotype or a distinct biological subtype with better clinical outcomes. To further elucidate this paradox, future studies should stratify patients by molecular subtypes or tumor stages, as recent spatial omics studies have revealed profound heterogeneity in the KIRP immune microenvironment [8]. It is plausible that LINC02471 expression marks a subtype with enhanced immune memory or altered stromal interactions, which could explain its association with both advanced pathology and favorable prognosis.
From a diagnostic perspective, LINC02471 demonstrated excellent discriminatory power between tumor and normal tissues, with an AUC of 0.887. This performance compares favorably with previously reported RNA-based biomarkers in RCC [26,27] and underscores its potential clinical utility. Given the increasing interest in non-invasive diagnostic strategies, such as circulating RNA detection, LINC02471 may represent a promising candidate for future liquid biopsy applications [28].
One of the most intriguing findings of this study is the significant negative correlation between LINC02471 expression and the infiltration of plasmacytoid dendritic cells (pDCs), activated dendritic cells (aDCs), and NK cells. These immune cell populations play critical roles in antigen presentation, innate immune surveillance, and the initiation of anti-tumor immune responses [29-31]. Reduced infiltration of these cells is often associated with impaired immune activation and immune escape. However, the favorable prognosis observed in patients with high LINC02471 expression suggests that its prognostic value may not be driven by enhanced innate immune cell recruitment.
Instead, LINC02471 may be involved in shaping a distinct immune microenvironment characterized by alternative immune regulatory mechanisms. For example, its positive correlation with eosinophils and central memory T cells (Tcm) hints at a potential role in modulating adaptive immune responses or immune memory formation, which have been associated with improved long- term cancer outcomes [32,33]. Alternatively, LINC02471 expression may reflect an intrinsic tumor biology that is less dependent on immune evasion mechanisms mediated by dendritic cells and NK cells.
Our in vitro experimental validation using multiple KIRP cell lines further strengthens the clinical relevance of LINC02471. The differential expression pattern observed across Caki-2, ACHN, and KTCL-140 cells mirrors the heterogeneity observed in patient samples, suggesting that LINC02471 expression may vary among different molecular subtypes of KIRP. The functional experiments demonstrating that LINC02471 knockdown inhibits proliferation while its overexpression promotes growth provide mechanistic insight into its role in KIRP progression. These findings are particularly significant given that Caki-2 cells, which exhibited the highest LINC02471 expression, are derived from a primary KIRP tumor and represent a clinically relevant model system.
Collectively, these findings highlight the complex and context-dependent role of LINC02471 in KIRP. While our study is limited by its retrospective nature and reliance on bioinformatic analyses, it provides a strong rationale for future mechanistic investigations. We are currently conducting in vitro and in vivo functional studies to validate the biological roles of LINC02471 in KIRP, including its effects on tumor invasion, migration, and immune modulation. The results from these ongoing experiments will be reported in a subsequent manuscript. Functional studies in vitro and in vivo will be essential to clarify how LINC02471 regulates tumor behavior and immune cell interactions, and whether it can be exploited as a therapeutic target or integrated into immune-based treatment strategies.
CONCLUSION
In summary, our integrative analysis identifies LINC02471 as a novel and pivotal molecule in KIRP. It functions as a highly sensitive diagnostic biomarker and an independent indicator of favorable prognosis. More importantly, its significant associations with specific immune cell populations illuminate its potential role in sculpting the KIRP immune landscape. These findings not only enhance our understanding of KIRP biology but also lay a foundational framework for future research. Validating LINC02471's function in vitro and in vivo, and exploring its utility in clinical settings as part of a multi-analyte signature, will be crucial steps toward translating this knowledge into improved patient stratification and novel therapeutic strategies for kidney renal papillary cell carcinoma.
ACKNOWLEDGMENTS
Not applicable.
FUNDING
Not applicable.
Availability of data and materials
Not applicable.
AUTHORS' CONTRIBUTIONS
All authors contributed to the study conception and design. All authors declared no competing interests. Dengwang Chen and Xinyue Jiang: Conceptualization, Methodology, Investigation, Writing - Original Draft, Writing - Review & Editing. (These authors contributed equally to this work.). Linna Wei, Dongmei Li and Zudi Meng: Supervision, Project administration, Resources, Writing - Review & Editing.
Ethics approval and consent to participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
REFERENCES
- Linehan WM, Ricketts CJ. The Cancer Genome Atlas of renal cell carcinoma: findings and clinical implications. Nature Reviews Urology. 2019;16(9):539-52. [Crossref] [Google Scholar] [PubMed]
- Moch H, Cubilla AL, Humphrey PA, Reuter VE, Ulbright TM. The 2016 WHO classification of tumours of the urinary system and male genital organs part A: renal, penile, and testicular tumours. European urology. 2016;70(1):93-105. [Crossref] [Google Scholar] [PubMed]
- Cancer Genome Atlas Research Network Analysis working group: Baylor College of Medicine Creighton Chad J. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature. 2013;499(7456):43-9. [Crossref] [Google Scholar] [PubMed]
- Li J, Gu A, Tang N, Zengin G, Li MY, et al. Patient‐derived xenograft models in pan‐cancer: From bench to clinic. Interdisciplinary Medicine. 2025;3(5):e20250016. [Crossref] [Google Scholar]
- Motzer RJ, Bander NH, Nanus DM. Renal-cell carcinoma. New England Journal of Medicine. 1996;335(12):865-75. [Crossref] [Google Scholar] [PubMed]
- Choueiri TK, Motzer RJ. Systemic therapy for metastatic renal-cell carcinoma. New England Journal of Medicine. 2017;376(4):354-66. [Crossref] [Google Scholar] [PubMed]
- Wu Y, Sun R, Ren S, Zengin G, Li M. Neuronal Reshaping of the Tumor Microenvironment in Tumorigenesis and Metastasis: Bench to Clinic. Medicine Advances. 2025;3(4):364-71. [Crossref] [Google Scholar]
- Li MY, Zhang Q, Zengin G, Guo Q, Sun W. Spatial omics: Deciphering heterogeneity in the tumor immune microenvironment and resistance to immunotherapy. Curr. Proteomics. 2025;22(5):100055. [Crossref] [Google Scholar]
- Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nature reviews genetics. 2016;17(1):47-62. [Crossref] [Google Scholar] [PubMed]
- Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer cell. 2016;29(4):452-63. [Crossref] [Google Scholar] [PubMed]
- Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer research. 2017 ;77(15):3965-81. [Crossref] [Google Scholar] [PubMed]
- Liu SJ, Dang HX, Lim DA, Feng FY, Maher CA. Long noncoding RNAs in cancer metastasis. Nature Reviews Cancer. 2021;21(7):446-60. [Crossref] [Google Scholar] [PubMed]
- Liu Y, Zhang H, Fang Y, Tang D, Luo Z. Non-coding RNAs in renal cell carcinoma: implications for drug resistance. Biomedicine & Pharmacotherapy. 2023;164:115001. [Google Scholar] [PubMed]
- Peng WX, Koirala P, Mo YY. LncRNA-mediated regulation of cell signaling in cancer. Oncogene. 2017;36(41):5661-7. [Crossref] [Google Scholar] [PubMed]
- Atianand MK, Caffrey DR, Fitzgerald KA. Immunobiology of long noncoding RNAs. Annual review of immunology. 2017;35(1):177-98. [Crossref] [Google Scholar] [PubMed]
- Chen YG, Satpathy AT, Chang HY. Gene regulation in the immune system by long noncoding RNAs. Nature immunology. 2017;18(9):962-72. [Crossref] [Google Scholar] [PubMed]
- Denaro N, Merlano MC, Lo Nigro C. Long noncoding RNA s as regulators of cancer immunity. Molecular oncology. 2019 ;13(1):61-73. [Crossref] [Google Scholar] [PubMed]
- Liu X, Li Y, Jiang X, Deng YE, Ma C, et al. Long non-coding RNA: Multiple effects on the differentiation, maturity and cell function of dendritic cells. Clinical Immunology. 2022;245:109167. [Crossref] [Google Scholar] [PubMed]
- Laskowski TJ, Biederstädt A, Rezvani K. Natural killer cells in antitumour adoptive cell immunotherapy. Nature Reviews Cancer. 2022;22(10):557-75. [Crossref] [Google Scholar] [PubMed]
- Erber J, Herndler-Brandstetter D. Regulation of T cell differentiation and function by long noncoding RNAs in homeostasis and cancer. Frontiers in Immunology. 2023;14:1181499. [Crossref] [Google Scholar] [PubMed]
- Chen D, Zhao H, Guo Z, Dong Z, Yu Y, et al. Identification of m6A-related lncRNAs LINC02471 and DOCK9-DT as potential biomarkers for thyroid cancer. International Immunopharmacology. 2024;133:112050. [Crossref] [Google Scholar] [PubMed]
- Mikami S, Oya M, Mizuno R, Kosaka T, Katsube KI, et al. Invasion and metastasis of renal cell carcinoma. Medical molecular morphology. 2014;47(2):63-7. [Crossref] [Google Scholar] [PubMed]
- Gutschner T, Diederichs S. The hallmarks of cancer: a long non-coding RNA point of view. RNA biology. 2012;9(6):703-19. [Crossref] [Google Scholar] [PubMed]
- Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. 2018;172(3):393-407. [Crossref] [Google Scholar] [PubMed]
- Nojima T, Proudfoot NJ. Mechanisms of lncRNA biogenesis as revealed by nascent transcriptomics. Nature reviews Molecular cell biology. 2022;23(6):389-406. [Crossref] [Google Scholar] [PubMed]
- Chehade CH, Agarwal N. Molecular subtypes as potential biomarkers in renal cell carcinoma. Cancer Cell. 2024;42(5):736-8. [Crossref] [Google Scholar] [PubMed]
- Xie J, Zhong Y, Chen R, Li G, Luo Y, et al. Serum long non‐coding RNA LINC00887 as a potential biomarker for diagnosis of renal cell carcinoma. FEBS Open Bio. 2020;10(9):1802-9. [Crossref] [Google Scholar] [PubMed]
- Wan JC, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nature Reviews Cancer. 2017;17(4):223-38. [Crossref] [Google Scholar] [PubMed]
- Villadangos JA, Young L. Antigen-presentation properties of plasmacytoid dendritic cells. 2008;29(3):352-61. [Crossref] [Google Scholar] [PubMed]
- Colonna M, Trinchieri G, Liu YJ. Plasmacytoid dendritic cells in immunity. Nature immunology. 2004;5(12):1219-26. [Crossref] [Google Scholar] [PubMed]
- Myers JA, Miller JS. Exploring the NK cell platform for cancer immunotherapy. Nature reviews Clinical oncology. 2021;18(2):85-100. [Crossref] [Google Scholar] [PubMed]
- Grisaru-Tal S, Jacobsen EA, Munitz A. Evolving role for eosinophils in cancer: from bench to bedside. Trends in Cancer. 2025 Jun 12. [Crossref] [Google Scholar] [PubMed]
- Gavil NV, Cheng K, Masopust D. Resident memory T cells and cancer. Immunity. 2024;57(8):1734-51. [Crossref] [Google Scholar] [PubMed]